The Australian Agronomist Magazine’s Spring 2019 issue features an article from Neutrog’s Microbiologist and R&D Manager, Dr. Uwe Stroeher. Titled ‘The Plant Microbiome’, Uwe discusses soil microbes which can significantly enhance plant health and growth, as well as positively affect crop yield in a world with a growing population and a potentially changing environment, where food security has become a major agricultural challenge.
In a world with a growing population and a potentially changing environment, food security has become a major agricultural challenge. We have known for many years that there is a range of soil microbes which can significantly enhance plant health and growth, as well as positively affect crop yield.
Unfortunately, until recently, the ability to decipher this plant-microbe interaction and to determine exactly which microbes are in the soil (or for that matter associated with the plant) has been difficult and extremely expensive. So why has it taken so long to be able to undertake these kinds of studies? There have been two major factors.
Firstly, traditional microbial studies are limited by the ability to grow various micro-organisms in a laboratory setting, so until now only ~2% of the potential soil bacteria and fungi can be cultured. We know even less about what they do, and how and when they are actually in the soil, or when they associate with plants.
Secondly, a major technical leap was required in order to examine the microbes using genomic sequencing. So we now find ourselves in a new age of meta genomics, where, thanks to Next Generation Sequencing, we have the ability to examine and piece together the whole range of DNA of micro-organisms that are found in the soil or associated with the plant. It is this association of plants with viruses, bacteria and fungi that constitutes the plant microbiome. Every tissue of the plant, whether it be leaves, roots or even seeds, interact in some way with various microbes, and in some cases micro-organisms (known as endophytes) are even internalised within plant tissue.
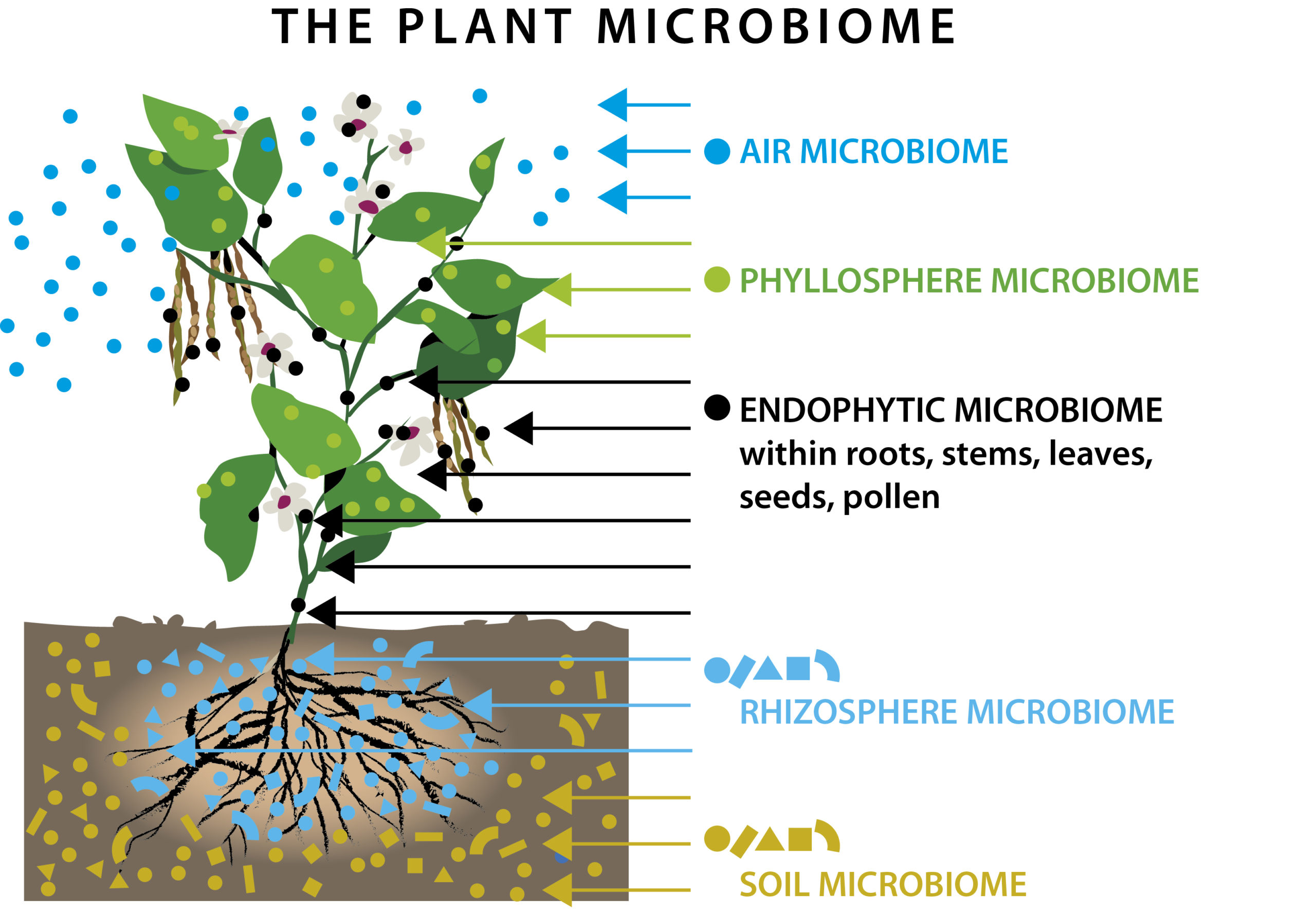
However, for ease of understanding, the plant microbiome is often split into two parts – the phyllosphere, which is the part above the ground – this describes the microbes associated with the stem, leaves, flowers, fruit and seeds of the plants. The second region is the rhizosphere, which represents the roots and the region surrounding the roots, where secretions from the plant can and will influence the microbes present in this area. (Fig 1)¹.
Over the last four to five years there has been an explosion of data on the plant microbiome. What we have learned is that the plant microbiome is likely to be the most diverse biome on the planet. As such, there is not a one-size-fits-all, although a core plant microbiome has been identified¹.
Therefore, every plant has its own unique set of micro-organisms, which is overlayed by the fact that the plant microbiome is not fixed, but is in continual flux. This flux or change can be due to environmental factors such as temperature, moisture, soil pH and even the nutritional status of the plant.
There are also means by which plants directly alter their microbiome. The most well studied is root secretions. In this way, plants can select and also discourage certain bacteria and fungi by secreting a range of sugars, organic acids, amino acids, phenolic compounds and plant growth factors – all of which alter the microbiome, in particular in the rhizosphere²⁻³.
Although we can now identify which microbes are present and no longer have to rely on classical microbiology, we still struggle to assign very specific functions to the many bacteria, viruses and fungi which co-habitate with plants. So what has the ability to determine plant microbiome really given us? We know that higher levels of diversity in the plant microbiome appears to be associated with better outcomes, and that the loss of even low represented species can be detrimental¹.
Many of the beneficial effects seen from the association of plants with micro-organisms can now be more readily mapped to an organism with specific characteristics or genes, such as the gene for 1-aminocyclopropane-1-decarboxylase, which is capable of reducing the level of the stress hormone ethylene, thereby reducing the damage done by this hormone.
It is now also possible to accurately map broad functional groups and genes involved in certain metabolic pathways, and to determine their relative predominance⁴. For example, if we see numerous genes involved in nitrogen fixation, or if we find many genes involved in the production of plant growth factors such as auxins or cytokinin, then it is most likely that these soils will effectively fix nitrogen and enhance plant growth respectively due to the microbes present.
Alternatively, if we see the presence of certain pathogenic genes such as those involved in toxin production, we can then speculate that these soils are more prone to allow disease-causing organism to flourish, or that plants in these soils are more likely to show disease. Conversely, it is now also possible to look for the biological control agents against microbes that may represent pathogens.
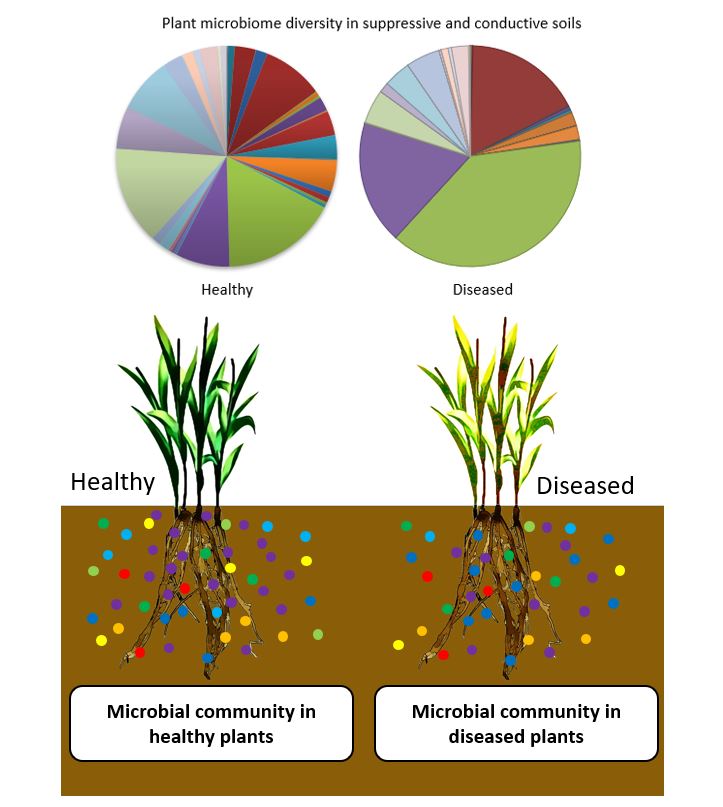
The other obvious advantage is that it’s now relatively easy to make a list of pathogens found, so if there is a dominance or high number of certain pathogens, this would highlight an issue prior to the disease showing up in the plants themselves (Fig2). Furthermore, the ability to unravel the plant microbiome has also allowed us to examine the difference between pathogen suppressive and permissive soils⁵. In suppressive soils, the pathogens are generally present in low numbers or have difficulty in establishing themselves. One characteristic feature of suppressive soils is that they have a higher diversity of microbes included in the rhizosphere⁶. A prime example is Take-All caused by the Gaeumannomyces graminis fungus, which, to some extent, can be controlled in suppressive soils by the build-up of fluorescent Pseudomonas spp⁶.
So what is the future for the plant microbiome? The ability to compare productive to non-productive soils at a microbial level will be possible, which may lead to the development of microbial inoculants (such as Neutrog’s GOGO Juice) to overcome issues in poorly performing soils. Additionally, the microbiome technology will subsequently allow the monitoring of the effect of any such inoculant on the plant’s microbiome, and whether these products can elevate issues¹.
It will also be possible to monitor the impact that certain cropping or growing practices have on the microbiome of the plants and soil, thereby allowing farmers to potentially mitigate any effects prior to crop losses or yield reductions.
Further down the track we will have the ability to select plants capable of interacting and taking advantage of beneficial microbes. We know from plant microbiome studies that domesticated plants have a lowered capacity to interact with plant growth-promoting micro-organisms⁷⁻⁸.
Neutrog are currently working together with Western Sydney University on a species of soil fungi called Trichoderma. Trichoderma is a known biological control agent which has the ability to effectively compete with plant pathogens, and can help to strengthen the overall defence of the plant. Neutrog have isolated a range of Trichoderma strains and are now testing their efficacy against a range of fungal plant pathogens on a number of economically important crops.
Journal References:
1. Compant, S. Samad, A. Faist, H. Sessitsch, A. A review on the plant microbiome: Ecology, function and emerging trends in microbial application. J Advanced Research. 2019; 19: 29-37. 2. Mendes, R. Garbeva, P. Raajmakers, JM. The rhizosphere microbiome: significance of plant beneficial plant pathogenic and human pathogenic microorganisms. FEMS Microbiol Rev. 2013; 37: 634-663 3. Fierer, N. Embracing the unknown: disentangling the complexities of the soil microbiome. Nature Rev. Micorbiol. 2017; 15: 579-590 4. Sergaki, C. Laguna, B. Lidbury, I. Gifford, ML. Schäfer P. Challenges and approaches in microbiome research: From fundamental to applied. Frontier in Plant Science 2018; 8: Article 1205 5. Schlatter, D. Kinkel, L. Thomashow, LS. Weller, DM. Paulitz, T. Disease suppressive soils: New insights from the soil microbiome. Phytopathol. 2017; 107: 1284-1297. 6. Weller, DM. Raaijmakers, JM. McSpadden Gardener, BB. Thomashow, LS. Microbial populations responsible for specific suppressiveness to plant pathogens. Annu. Rev. Phytopathol. 2002; 40: 309-348. 7. Pérez-Jaramillo, JE. Mendes, R. Raaijmakers JM. Impact of plant domestication on rhizosphere microbiomes assembly and functions. Plant Mol. Biol. 2016; 90: 635-644. 8. Gopal, M. Gupta, A. Microbiome selection could spur next-generation plant breeding strategies. Frontier in Plant Science 2016; 7: Article 1971
